 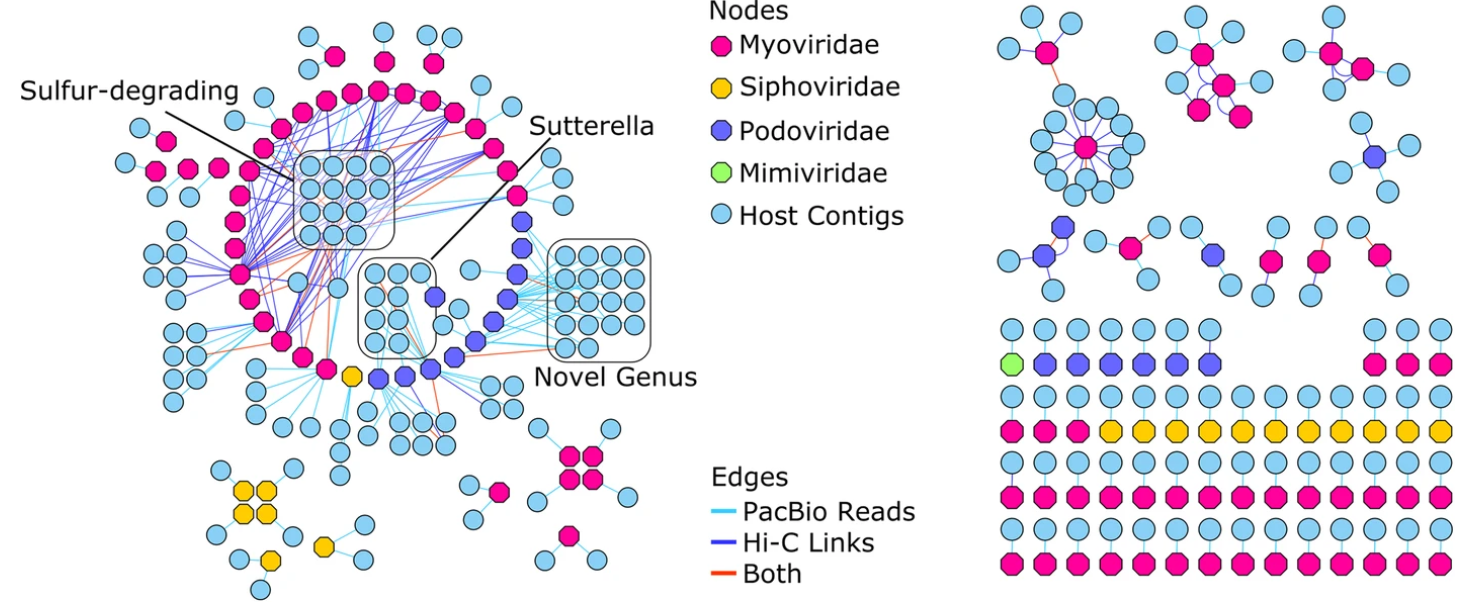 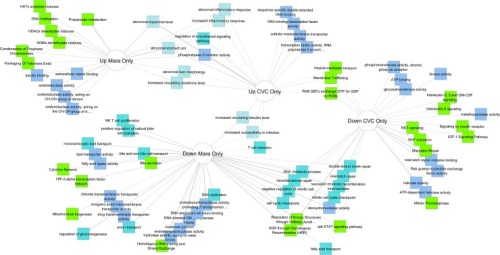
Furthermore, IFI27 was confirmed to be the most significantly expressed in clinical verification. Subsequently, 13 hub genes were identified, and function and pathway enrichments of hub genes mainly were: response to virus, defense response to virus, regulation of viral genome replication and regulation of viral life cycle. Totally, 4028 DEGs were screened out and among which, 131 most significant DEGs were selected. Finally, the gene set enrichment analysis (GSEA) focusing on IFI27 was carried out. Further, the RSV infected infants with high-expression IFI27 and those with low-expression IFI27 were compared (defined as higher or lower than the median mRNA level). CYTOSCAPE VIRAL VERIFICATIONClinical verification was employed to verify the expression level of top five hub genes among 72 infants including 50 RSV infected patients and 22 non-RSV-infected patients hospitalized in our center. CYTOSCAPE VIRAL SOFTWAREThe module analysis was performed by Cytoscape software and hub genes were identified. The protein-protein interaction (PPI) network for DEGs was constructed through Search Tool for the Retrieval of Interacting Genes (STRING). The function and pathway enrichments were analyzed through Database for Annotation, Visualization and Integrated Discovery (DAVID) website. Three datasets (GSE77087, GSE69606 and GSE41374) containing 183 blood samples of RSV infected patients and 33 blood samples of healthy controls from Gene Expression Omnibus (GEO) database were downloaded and the differentially expressed genes (DEGs) were screened out. The purpose of the present study was to identify candidate genes of preterm infants who are susceptible to RSV infection and provide a new insight into the pathogenesis of RSV infection. Cytoscape is being extended through apps and core features to search, extract, visualize, and analyze data from the Human Cell Atlas, with a focus on network and pathway analysis.Preterm infants are a special population that vulnerable to respiratory syncytial virus (RSV) infection and the lower respiratory tract infections (LRTIs) caused by RSV could be severe and even life-threating. Most of the Apps are freely available from Cytoscape App Store. Apps are available for network and molecular profiling analyses, new layouts, additional file format support, scripting, and connection with databases.Īpps may be developed by anyone using the Cytoscape open API based on Java™ technology and App community development is encouraged. Additional features are available as Apps (formerly called Plugins). Although Cytoscape was originally designed for biological research, now it is a general platform for complex network analysis and visualization.Ĭytoscape's core distribution provides a basic set of features for data integration, analysis, and visualization. Cytoscape is an open source software platform for visualizing molecular interaction networks and biological pathways and integrating these networks with annotations, gene expression profiles and other state data. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
